Detection of some indels was challenging for the routine IonReporter ™ workflow however, the addition of NextGENe® software improved indel identification demonstrating the importance of both bench and bioinformatic validation. We observed a good sequencing performance based upon amplicon, gene (hotspot variants within gene) and sample specific analysis with 92% of clinical samples obtaining an average amplicon coverage above 500X. The Oncomine ™ Focus Fusion panel had a good sensitivity and specificity based upon the samples assessed, however requires further validation to confirm findings due to limited sample numbers. The Oncomine ™ Focus DNA assay had a sample and variant-based sensitivity of 99.1 and 97.1%, respectively, and an assay specificity of 100%. This validation considered a number of parameters including the clinical robustness of the bioinformatics pipeline for variant detection and interpretation. 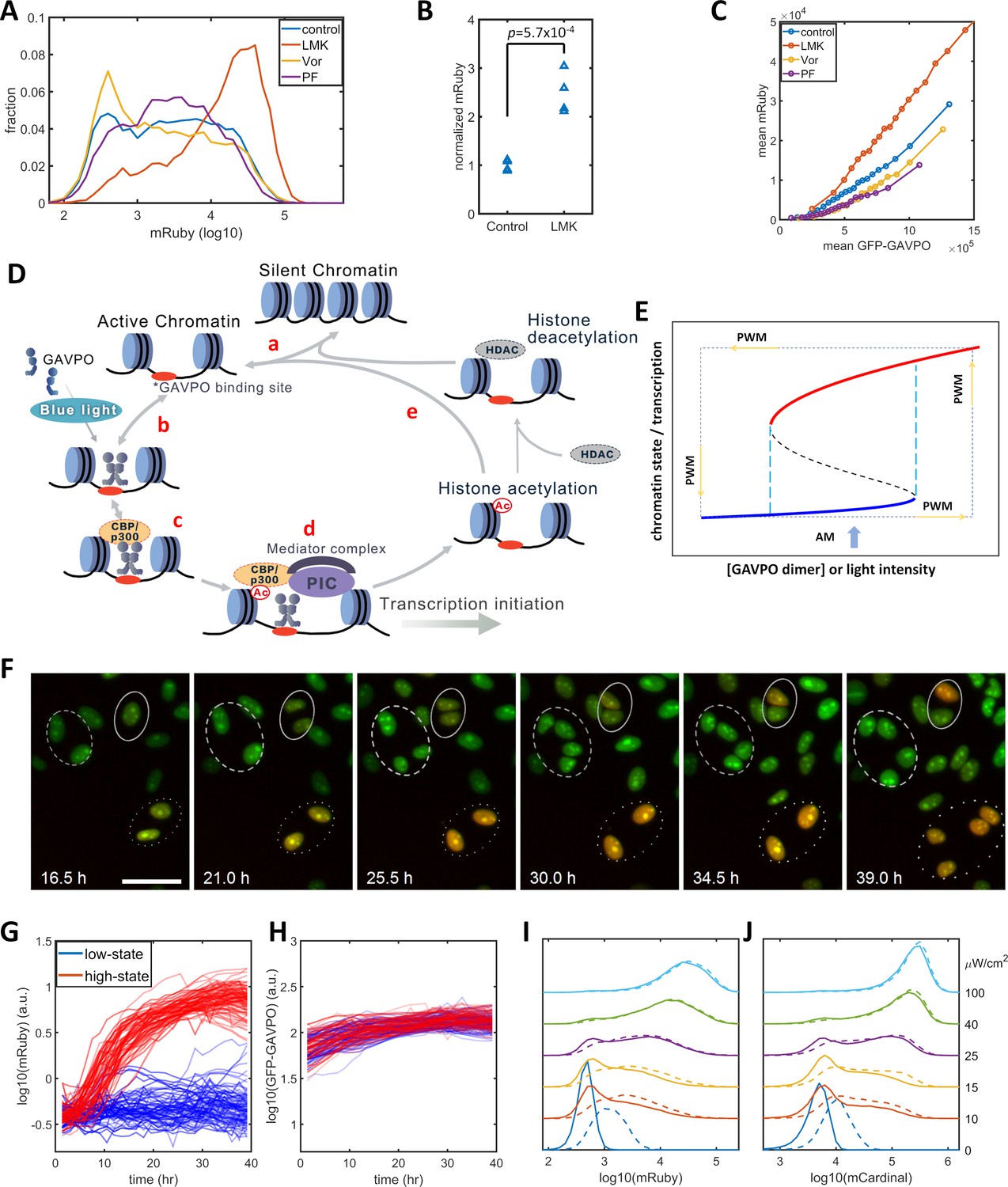
Using a mixture of routine FFPE and reference material across a variety of tissue and specimen types, we sequenced 86 and 31 samples on the Oncomine ™ Focus DNA and RNA Fusion assays, respectively. The aim of this clinical validation was to assess the performance of the Ion Torrent Personal Genome Machine (IonPGM ™) and validate the Oncomine ™ Focus DNA and RNA Fusion panels for clinical application in solid tumour testing of formalin-fixed, paraffin-embedded (FFPE) tissue. However, translation into daily use requires a rigorous and comprehensive validation strategy. The clinical utility of next-generation sequencing (NGS) for a diverse range of targets is expanding, increasing the need for multiplexed analysis of both DNA and RNA. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
